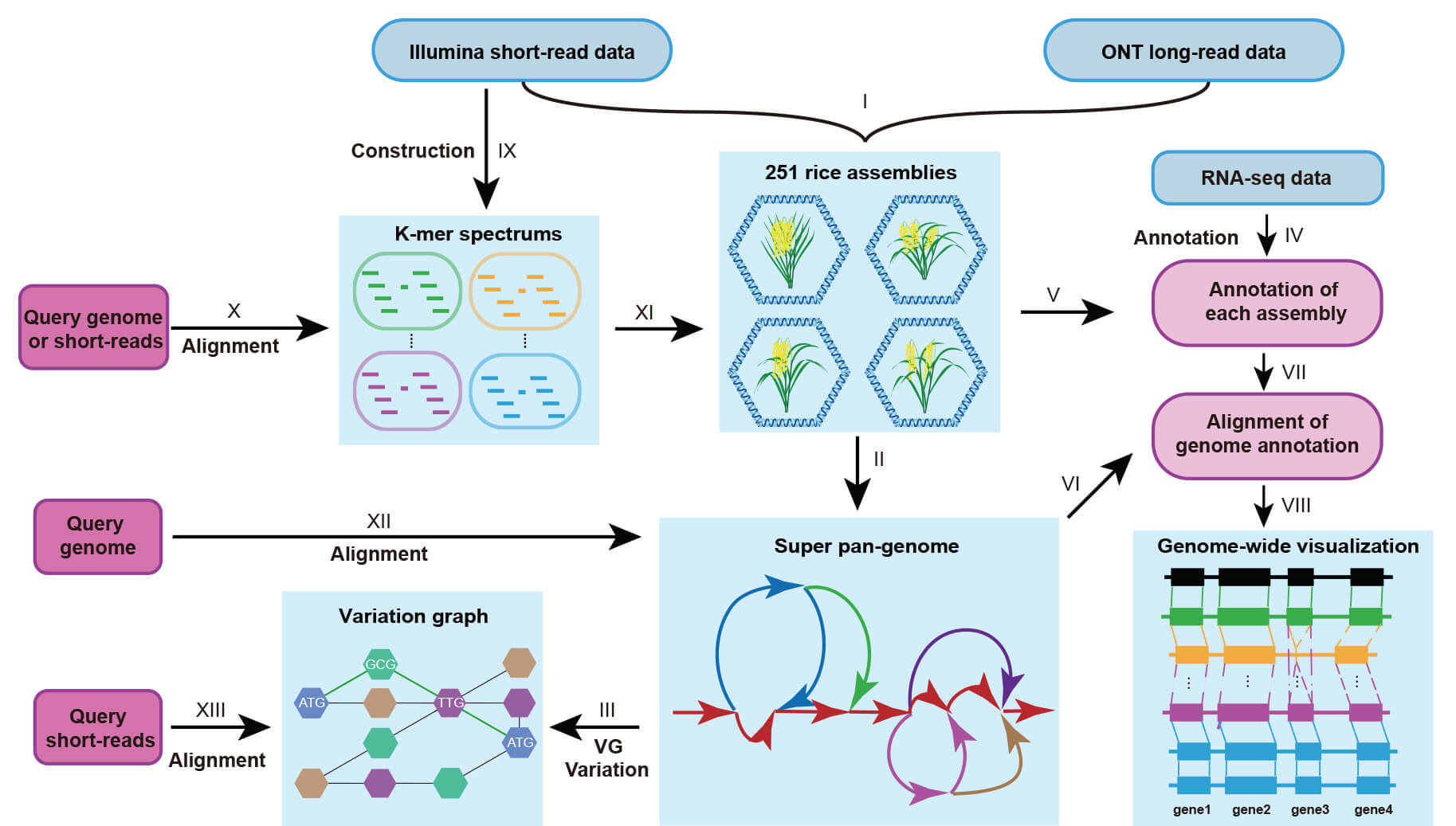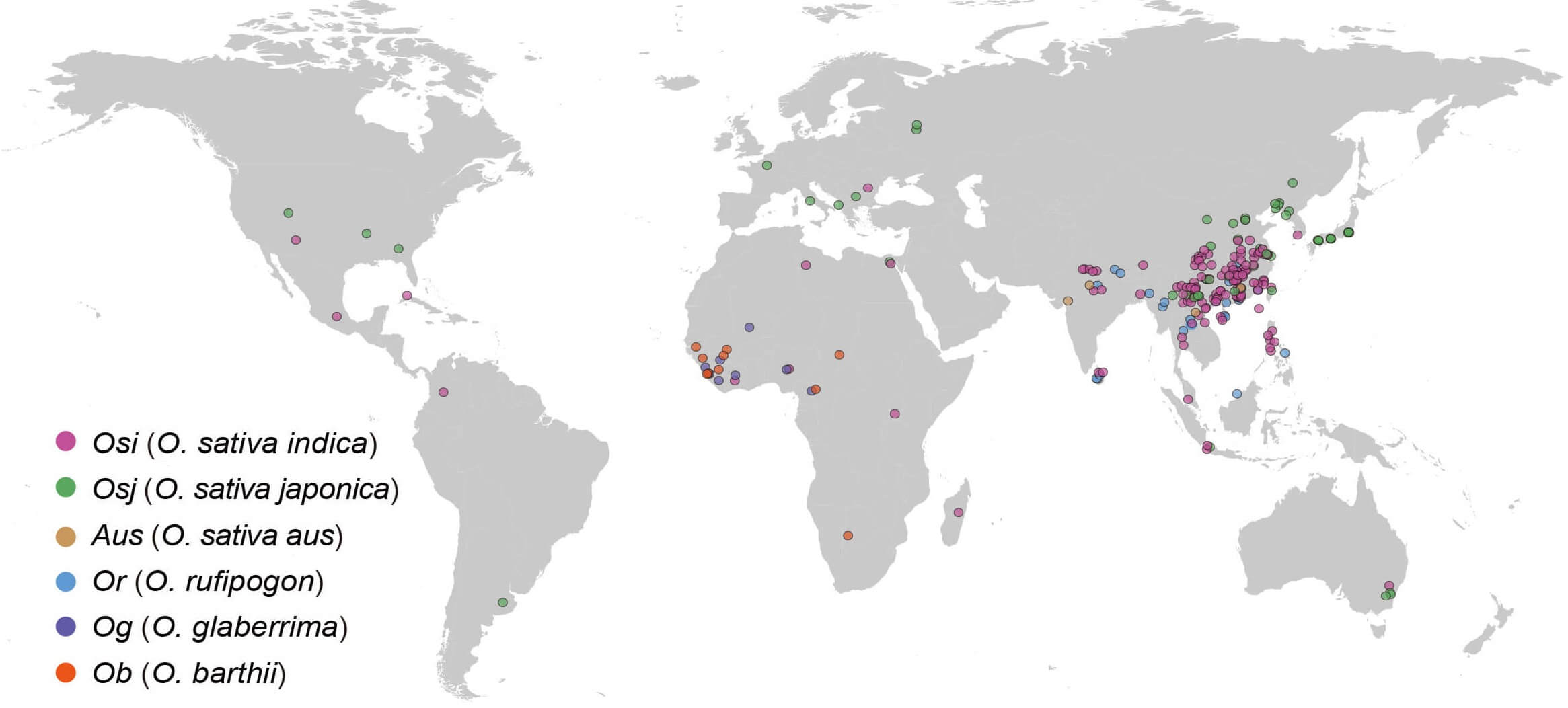
The super pan-genome of Wild & Cultivated Rice Project based on reference-quality de novo long-read assemblies (202 O. sativa, 28 O. rufipogon, 11 O. glaberrima, and 10 O. barthii) accessions aimed to generate a super pan-genome and super pan-annotation. To facilitate the applications of these results, we developed the Rice Super Pan-genome Information Resource Database, an interactive web-based browser for the rice super pan-genome. The reference-free whole-genomes alignment of 251 accessions, genome annotation, pan-genome graph, pan-genome variations, K-mer spectrums and a variation graph based on pan-genome variations were also included in the RiceSuperPIRdb. Besides, more data and analysis results will also be included in our database soon.

1.Basic information of the 251 rice accessions, including the genome sequence, gene annotations, transposable element annotations, and structural variation;
2.Reference-free multiple assembly alignment, which means that the user can choose any accession as a reference to browse whole genome alignment;
3.Pan-genome and annotation information, including a graph-based pan-genome of rice accessions by minigraph. Gene annotation and transposable elements of each assembly were associated with the pan-genome. A variation graph using the variations from the pan-genome could be downloaded, which could be used to link query short-reads to the super pan-genome variations.
4.Node-specific k-mer spectrums for each of 256 accessions, including our 251 rice accessions and 5 public accessions (O.punctata, O.meridionalis, O.glumaepatula, Nipponbare, and R498), which can be used to verify their closest assembly reference with low-quality short-read data.